CLC Assembly Cell is constantly under development. Does anyone have experience in generating workflow in CLC Genomics Workbench 7. Download CLC Genomics Workbench 8.5.1 Full + crack Download clc genomics workbench v.5.5.2 crack elite. Clc Genomics Workbench 5.1 Crack - eraditceive For enhancing functionalities of CLC Genomics Workbench. BMC Genomics 15:15īolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. The Workbench Modules is intended for molecular biology applications. Hou CY, Wu MT, Lu SH, Hsing YI, Chen HM (2014) Beyond cleaved small RNA targets: unraveling the complexity of plant RNA degradome data. You can create and edit alignments, work with interactive restriction site analysis, phylogenetics, or. CLC Sequence Viewer includes a number of features for doing basic bioinformatics analysis. Lin PC, Lu CW, Shen BN, Lee GZ, Bowman JL, Arteaga-Vazquez MA, Liu LY, Hong SF, Lo CF, Su GM, Kohchi T, Ishizaki K, Zachgo S, Althoff F, Takenaka M, Yamato KT, Lin SS (2016) Identification of miRNAs and their targets in the liverwort Marchantia polymorpha by integrating RNA-Seq and degradome analyses. Download popular programs, drivers and latest updates easily. Many degradome reads map to the gene sequence. 11 In this training, we’ll use QIAGEN CLC Genomics. Webinar: Pathogen detection and variant analysis using hybrid capture technology Jul. 
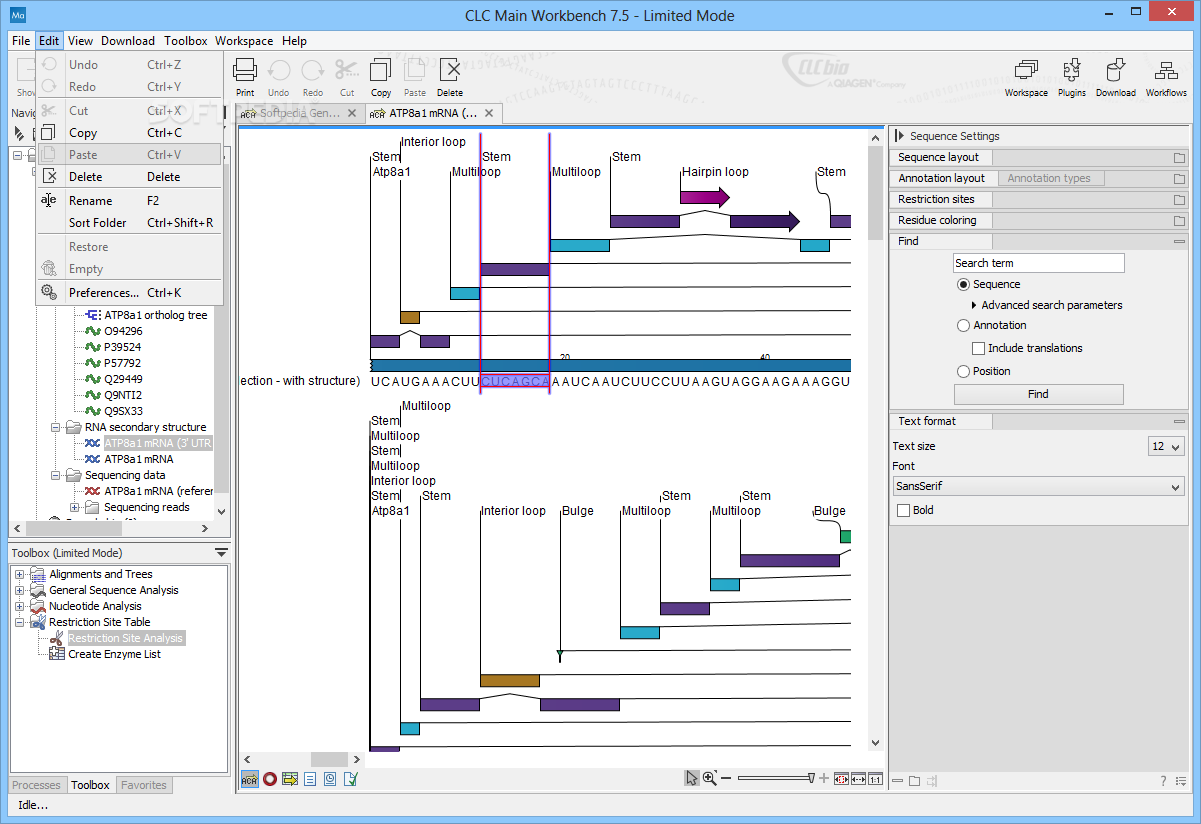
11 Join Leonie Hodges, Scientific Communications Officer for COSMIC, for this webinar to take a journey through COSMIC’s mission. We use the GROWTH-REGULATING FACTOR1 gene (GRF1 AT2G22840) as an example, and the mapping results are shown in Fig. Webinar: Insights into cancer genomics via COSMIC v98 Jul. Zhai J, Arikit S, Simon SA, Kingham BF, Meyers BC (2014) Rapid construction of parallel analysis of RNA end (PARE) libraries for Illumina sequencing. The mapping results can be viewed through the CLC Genomics Workbench interface (Fig. Niu QW, Lin SS, Reyes JL, Chen KC, Wu HW, Yeh SD, Chua NH (2006) Expression of artificial microRNAs in transgenic Arabidopsis thaliana confers virus resistance. Souret FF, Kastenmayer JP, Green PJ (2004) AtXRN4 degrades mRNA in Arabidopsis and its substrates include selected miRNA targets. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |



 RSS Feed
RSS Feed
